CNeuroMod 2020-alpha
CNeuroMod makes its first internal release!
Wed, Feb 5, 2020 By Pierre Bellec.
Concretely, this means that a whole pipeline for data production is in place, and that the team is now running analyses on CNeuroMod data for the first time. This post goes through the main steps of the “data release” pipeline, and explains which tools we chose for completing those steps. Huge props to all the people who have helped build these fantastic tools, and in particular Michael Hanke and the studyforest team, which have been a huge inspiration. This is euphemism to say we copied or reused most of what they have done…
 The first step of the pipeline is converting all the data in a standardized, easy-to-reuse and self-documented organization. We chose the Brain Imaging Data Standard (BIDS) format, because, well, it’s the only one. At this stage, the release does not include all the physiological signals (cardiac, respiration, etc), or the responses of the subject during the
The first step of the pipeline is converting all the data in a standardized, easy-to-reuse and self-documented organization. We chose the Brain Imaging Data Standard (BIDS) format, because, well, it’s the only one. At this stage, the release does not include all the physiological signals (cardiac, respiration, etc), or the responses of the subject during the hcptrt tasks. These will come in due time, and represent another big chunk of work. But the release does include all the stimuli, with edited movie segments of movie10 and a tsv description of the events presented to the subjects during the tasks of hcptrt. Note that all stimuli presentation scripts are also available on our CNeuroMod tast_stimuli github repository.
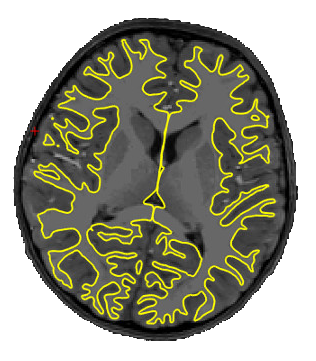 We know that the neuroimaging community loves preprocessing data, and will likely want to use their trusted in-house approach starting with the raw data. We still wanted to have a reference preprocessing version available as a point of comparison. There are also people out there who will be interested in using preprocessed data, either because they are lazy (most people in our team share this great quality) or simply lack the domain specific knowledge or computational resources to run this processing. So, as part of each release, we will include a number of processed derivatives. This first
We know that the neuroimaging community loves preprocessing data, and will likely want to use their trusted in-house approach starting with the raw data. We still wanted to have a reference preprocessing version available as a point of comparison. There are also people out there who will be interested in using preprocessed data, either because they are lazy (most people in our team share this great quality) or simply lack the domain specific knowledge or computational resources to run this processing. So, as part of each release, we will include a number of processed derivatives. This first 2020-alpha release includes the outputs of the fMRIprep pipeline, along with the brain surface reconstruction results generated with Freesurfer. More derivatives, including quantitative MRI metrics and high-resolution individual functional parcellations, will soon follow.
 The next step of the pipeline is a DataLad instance. DataLad is basically a git repository, with all the file names and the meta-data (text files, json files). But the files themselves are just pointers to a S3 bucket, and are not accessible with just a clone of the DataLad. To effectively download the files through the S3 bucket, a login and password are required. We are setting up a form to request access and login credentials. At this stage the DataLad instance is private, hosted on our server, but we will push a fork to our public github page at each public release. This way we can have issue tracking and “live” update of the database internally, and at the same time share publicly all released data structures. With this approach, individuals will be able to browse the file structure and all meta data on github, without requesting access. The access form will only be required to effectively download the data files through S3.
The next step of the pipeline is a DataLad instance. DataLad is basically a git repository, with all the file names and the meta-data (text files, json files). But the files themselves are just pointers to a S3 bucket, and are not accessible with just a clone of the DataLad. To effectively download the files through the S3 bucket, a login and password are required. We are setting up a form to request access and login credentials. At this stage the DataLad instance is private, hosted on our server, but we will push a fork to our public github page at each public release. This way we can have issue tracking and “live” update of the database internally, and at the same time share publicly all released data structures. With this approach, individuals will be able to browse the file structure and all meta data on github, without requesting access. The access form will only be required to effectively download the data files through S3.
 The final step of our “data release” pipeline is a pretty detailed documentation, which can be found at docs.cneuromod.ca. The raw content is public, and hosted on a github repo. Because CNeuroMod will regularly release new data, with new features, we basically treated that data project like software. The documentation is released with version numbers that match the ones used for the DataLad releases (more on that latter). Any change pushed to the master branch of the documentation triggers an automatic rebuild of the documentation website, thanks to the awesome service provided by read-the-docs. If a reader clicks the small box at the bottom left corner of the website (on desktops), it is possible to access any release of the documentation, as well as download the documentation as a
The final step of our “data release” pipeline is a pretty detailed documentation, which can be found at docs.cneuromod.ca. The raw content is public, and hosted on a github repo. Because CNeuroMod will regularly release new data, with new features, we basically treated that data project like software. The documentation is released with version numbers that match the ones used for the DataLad releases (more on that latter). Any change pushed to the master branch of the documentation triggers an automatic rebuild of the documentation website, thanks to the awesome service provided by read-the-docs. If a reader clicks the small box at the bottom left corner of the website (on desktops), it is possible to access any release of the documentation, as well as download the documentation as a pdf or epub document! In reality, we are thinking about this documentation much more like a living book than a website.
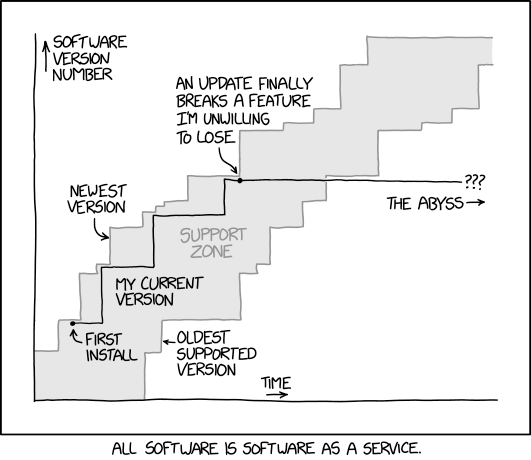 One last important point is how we name our releases, and how often we plan to release data. We decided to aim for one release per year, which should represent roughly 100 hours of fresh functional neuroimaging data per subject. Each release is simply named after the year, so we are currently working towards the
One last important point is how we name our releases, and how often we plan to release data. We decided to aim for one release per year, which should represent roughly 100 hours of fresh functional neuroimaging data per subject. Each release is simply named after the year, so we are currently working towards the 2020 release. But each of these official releases will be preceded by three “pre-release”. The YEAR-alpha release is done early in the release cycle and kept internal to the CNeuroMod team. It gives us an opportunity to start analyses on recently collected data, and check that they work as expected. The YEAR-beta release includes all the data planned for the official release, but has only been looked at by few people. We share these data with an extended pool of CNeuroMod beta testers, who are colleagues directly collaborating with us and agreeing to report minor issues. Finally the YEAR-rc release is our candidate release, which we think is basically ready to go. We will still wait a few months to collect feedback before freezing an official release. During the Human Brain Mapping conference in June 2020, Montreal, we will announce our first public release, 2020-rc. We are aiming for the 2020 release in September 2020.
Congrats for reading through this long and possibly unnecessarily detailed first blog post! Here’s some random ASCII art as a reward!
_________ __ .__
\_ ___ \ ____ __ __________/ |_ ____ |__| ______
/ \ \/ / _ \| | \_ __ \ __\/ _ \| |/ ___/
\ \___( <_> ) | /| | \/| | ( <_> ) |\___ \
\______ /\____/|____/ |__| |__| \____/|__/____ >
\/ \/
_______ .___
\ \ ____ __ _________ ____ _____ ____ __| _/
/ | \_/ __ \| | \_ __ \/ _ \ / \ / _ \ / __ |
/ | \ ___/| | /| | \( <_> ) Y Y ( <_> ) /_/ |
\____|__ /\___ >____/ |__| \____/|__|_| /\____/\____ |
\/ \/ \/ \/
